Research

Immunological Mandala: a metaphor for the universe of immune components in the body.

Integrative Immunology Laboratory
If the immune system was an unknown language to decode, then we likely know most letters (genes) and words (cell types), but it remains difficult to make sentences (cell-cell communications), let alone write coherent paragraphs (tissue- and organism-level processes). This illiteracy is perhaps best exemplified by our inability to manipulate the immune system against many deadly diseases and highlights the need to develop new ways to observe, quantify and ultimately decode immunity.
Our research aims to uncover the grammar – or general rules – governing how successful immune responses work to stop an infection or remove diseased cells while keeping the host healthy at the end of the process. We emphasize a multiscale approach that parallels the organization of the immune system across the body from molecules and cells to tissues to the whole organism. By creating new tools and approaches, we study how interactions occur between immunological components across these scales to uncover fundamental concepts in immunology and beyond. We pursue this goal in the hope that it will contribute knowledge to the longstanding question of how to manipulate the immune system at will.
Projects
Decoding the Body Language of Immunity
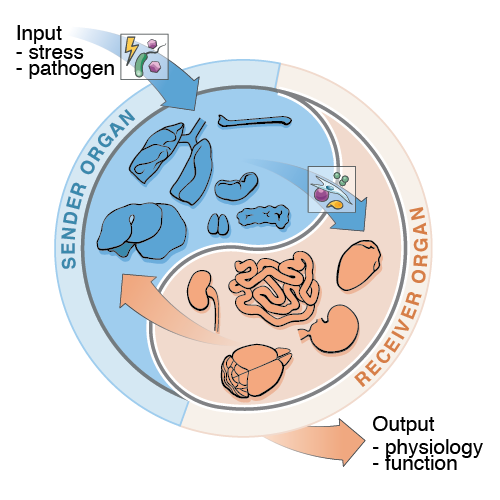
A fundamental challenge in immunology today is to develop new ways to study the structure, regulation, and function of the immunological events that cross organ boundaries. Deciphering the rules of interorgan immune signaling will yield insights into the functions and malfunctions of immunity at an unprecedented scale, that of the whole organism. To address this challenge, we recently developed tools for organism-wide expression profiling that led to the discovery of new interorgan mechanisms of host protection mediated by immune molecules and cells. We are working on applying these tools in comparative models of infectious and non-infectious diseases in rodents, which will: (1) help build a new field focused on the organism-wide probing of biological systems in mammals, (2) reveal new interorgan pathways that are crucial for immunity, and (3) contribute to our understanding of the functions and malfunctions of interconnected organ systems.
Takahama et al., Nature Immunology, 2024
Chevrier, Current Opinion in Systems Biology, 2019
Kadoki et al., Cell, 2017
The Combinatorics of Immune Signaling
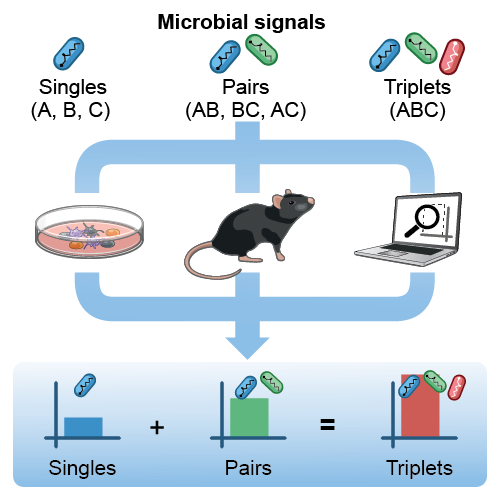
Biological systems make decisions in response to combinations of multiple signals. For example, to stop an infection, the immune system has learned to recognize and exploit the inter-dependencies of microbial signals by evolving in a chance-driven world of encounters with pathogens. We study how immune cells respond to combinations of multiple signals such as microbial molecules. In recent work, we discovered a general property for the combinatorial sensing of microbial signals, whereby the effects of triplet combinations of agonists on immune responses can be predicted by the effects of single and pairwise stimulations – using both cell cocultures in vitro and mouse models. Our finding greatly simplifies the description of the combinatorial problem posed by the sensing of complex microbial or adjuvant inputs by innate immune pathways and their downstream impact on immunity. We are building upon these results to (1) study the mechanisms of action of adjuvant combinations, and (2) study the fundamental rules of combinatorial sensing in various immune cells and pathways.
Immunotherapy & Vaccine Design
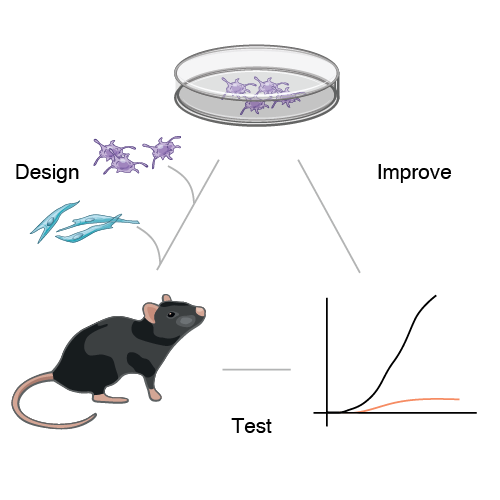
Live vaccines that are empirically attenuated from pathogens have been a powerful means to yield life-long immunity against many deadly pathogens. However, the rational design of non-live vaccines using immunomodulatory agents such as adjuvants has remained an elusive task in many cases where live vaccination is not efficacious or feasible. To tackle this challenge, a central question to answer is how do complex combinations of microbial or adjuvant signals shape immune responses? Filling this fundamental gap in our knowledge is critical to learn how to rationally choose and combine adjuvants to manipulate immunity against infectious and non-infectious diseases such as cancer. We discovered an intrinsic property governing the combinatorial logic of microbial sensing: the effects of triplet combinations of microbial signals can be accurately predicted using the data from the effects of singles and pairs of stimuli. In recent work, we exploited this simplifying property to develop and characterize vaccines using triplet combinations of adjuvants that led to protective immunity in mouse models of cancer. We are building upon these results to (1) identify new combinations of immunomodulatory agents useful for therapy, and (2) create new classes of immunomodulatory molecules.
Technology Development
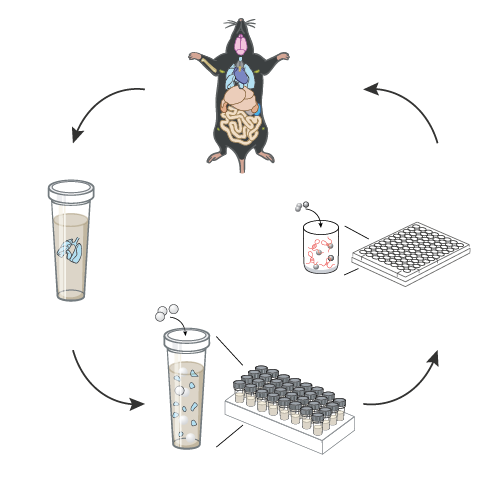
Immune cells are present in every organ of the body and every cell in the body carries immunological functions. To understand the rules of interdependency between immune cells and processes across multiple organs, it is crucial to study immune responses at the scale of the entire organism. However, we lack experimental approaches to systematically track the organism-wide processes that take place during immunological or other physiological responses in mammals. We aim to help build a new field focused on the probing of biological systems at the scale of the whole organism in mammals. For example, in recent work, we reported a suite of methods for the quantification of gene expression across most mouse organs. We demonstrated its use for the study of the molecular and cellular communication conduits that take place between organs during immune responses to vaccination and infection.